Premiering Monday, 4/22 on PBS NC
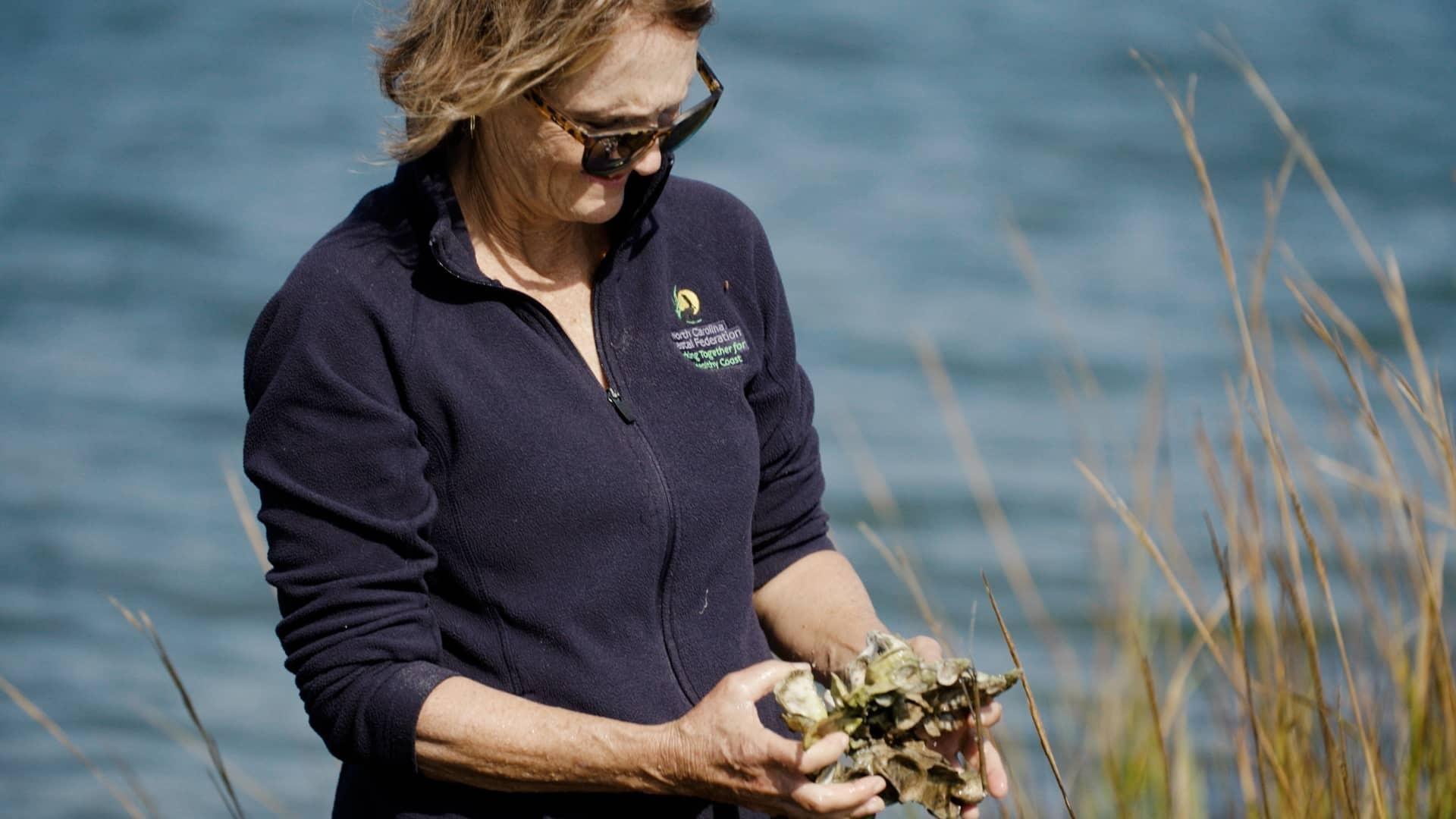
State of Change: Seeds of Hope
Meet the North Carolinians who are adapting to our changing climate in new and inspiring ways. From conserving land and growing food that’s tolerant of extreme conditions to using farmland to produce both crops and solar energy, these innovators are facing the challenges of climate change with perseverance and ingenuity.